So once more I find myself blogging on the subject of PAINS although in the wake of the 2015 Nobel Prize for medicine which will have forced many drug-likeness 'experts' onto the back foot. This time the focus is on antifungal research and the article in question has already been reviewed In The Pipeline. It's probably a good idea to restate my position on PAINS so that I don't get accused of being a Luddite who willfully disregards evidence (I'm happy to be called a heretic who willfully disregards sermons on the morality of ligand efficiency metrics and the evils of literature pollution). Briefly, I am well aware that not all output from biological assays smells of roses although I have suggested that deconvolution of fact from opinion is not always as quite as straightforward as some of the 'experts' and 'thought leaders' would have you believe. This three-part series of blog posts ( 1 | 2 | 3 ) should make my position clear but I'll try to make this post as self-contained as possible so you don't have to dig around there too much.
So it's a familiar story in
antifungals. More publications but less products and the authors of the featured article
note, “However, we believe that one key reason is the meager
quality of some of these new inhibitors reported in the antifungal
literature; many of which contain undesirable features” and I'd
have to agree that if the features really do cause compounds to choke
in (or before) development then the features
are can legitimately be described as 'undesirable'. As I've said before, it is one thing to opine that something looks yucky but another thing entirely to establish that it
really is yucky. In other words, it is not easy to deconvolute fact
from opinion. The authors claim, “It is therefore not surprising
to see a long list of papers reporting molecules that are
fungicidally active by virtue of some embedded undesirable feature.
On the basis of our survey and analysis of the antifungal literature
over the past 5 years, we estimate that those publications could
cover up to 80% of the new molecules reported to have an antifungal
effect”. I have a couple of points to make here. Firstly, it
would be helpful to know what proportion of that 80% have embedded
undesirable features that are actually linked to relevant bad behavior in
experimental studies. Secondly, this is a chemistry journal so
please say 'compound' when you mean 'compound' because 80% of even just a
mole of molecules is a shit-load of molecules.
So it's now time to say something
about PAINS but I first need to make the point that embedded
undesirable features can lead to different types of bad behavior by
compounds. Firstly, the compound can interact with assay components
other than target protein and I'll term this 'assay interference'. In
this situation, you can't believe 'activity' detected in the assay but you may be able to
circumvent the problem by using an orthogonal assay. Sometimes you can assess
the seriousness of the problem and even correct for it as described
in this article. A second type of bad behaviour is observed when the
compound does something unpleasant to the target (and other proteins)
and I'd include aggregators and redox cyclers in this class along with compounds that form covalent bonds with the protein in an unselective manner. The
third type of bad behaviour is
observed when the embedded undesirable feature causes the compound to
be rapidly metabolized or otherwise 'ADME-challenged'. This is the sort of bad behaviour that lead optimization teams have to deal with but I'll not be discussing it in this post.
The authors note that the original
PAINS definitions were derived from observation of “structural
features of frequent hitters from six different and independent
assays”. I have to admit to wondering what is meant by
'different and independent' in this context and it comes across as rather defensive. I have three questions for the authors. Firstly, if you
had the output of 40+ screens available, would you consider the
selection of six AlphaScreen assays to be an optimal design for an experiment to detect and characterize pan-assay interference?
Secondly, are you confident that singlet oxygen quenching/scavenging
by compounds can be neglected as an interference mechanism for these
'six different and independent AlphaScreen assays'? Thirdly, how many
compounds identified as PAINS in the AlphaScreen assays were shown to
bind covalently to one or more of their targets in the original PAINS article?
The following extract from the article
should illustrate some of difficulties involved in deconvoluting fact
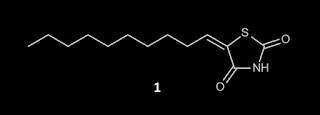
from opinion on the PAINS literature and I've annotated it in red. Here is the structure of compound 1:
"One class of molecules often reported
as antifungal are rhodanines and molecules containing related
scaffolds. These molecules are attractive, since they are easily
prepared in two chemical steps. An example from the recent patent
literature discloses (Z)-5-decylidenethiazolidine-2,4-dione
(1) [It's not actually a rhodanine and don't even think about bringing up the fact that my friends at Practical Fragments have denounced both TZDs and thiohydantoins as rhodanines] as a good antifungal against
Candida albicans (Figure2).(16)
As a potential carboxylic acid isostere, thiazolidine-2,4-diones may
be sufficiently unreactive such that they can progress some way in
development.(17) Nevertheless, one
should be aware of the thiol reactivity associated with this type of
molecule, as highlighted by ALARM NMR and glutathione assays,(18)
[I can't access reference 18 but rhodanines appear to represent its primary focus. Are you able to present evidence for
thiol reactivity for compound 1? What is the pKa for compound 1 and how might this be relevant to its ability to function as a Michael acceptor?] and this is especially relevant
when the compound contains an exocyclic alkene such as in 1.
The fact that rhodanines are promiscuous compounds has been recently
highlighted in the literature.(13, 18) [Maybe rhodanines really are promiscuous but, at the risk of appearing repetitious, 1,
is not a rhodanine and when we start extrapolating observations made
for rhodanines to other structural types, we're polluting fact with
opinion. Also the evidence for promiscuity presented in reference 13
is frequent-hitter behavior in a panel of six AlphaScreen assays and
this can't be invoked as evidence for thiol reactivity because rhodanines lacking the exocycylic carbon-carbon double bond are
associated with even greater PAIN levels than rhodanines that can
function as Michael acceptors] The observed antifungal activity of 1
is certainly genuine; however, this could potentially be the result
of in vivo promiscuous reactivity in which case the main issue lies
in the potential lack of selectivity between the fungi and the other
exposed living organisms" [This does come across as arm-waving. Have you got any evidence that there really is an issue? It's also worth remembering that, once you move away from the AlphaScreen technology, rhodanines and their cousins are not the psychopathic literature-polluters of legend and it is important for PAINS advocates to demonstrate awareness of this article.]
The authors present chemical structures of thirteen other compounds that they find unwholesome and I certainly wouldn't be be volunteering to optimize any of the compounds presented in this article. As an aside, when projects are handed over at transition time, it is instructive to observe how perceptions of compound quality differ between those trying to deliver a project and those charged with accepting it. My main criticism of the article is that very little evidence that bad things are actually happening is presented. The authors seem to be of the view that reactivity towards thiols is necessarily a Bad Thing. While formation of covalent bonds between ligands and proteins may be frowned upon in Switzerland (at least on Sundays), it remains a perfectly acceptable way to tame errant targets. A couple of quinones also appear in the rogues gallery and it needs to be pointed out that the naphthaquinone atovaquone is one of the two components (the other is proguanil which would also cause many compound quality 'experts' to spit feathers if they had failed to recognize it as an approved drug) of the antimalarial drug Malarone. I have actually taken Malarone on several occasions and the 'quinone-ness' of atovaquone worries me a great deal less than the potential neuropsychiatric effects of the (arguably) more drug-like mefloquine that is a potential alternative. My reaction to a 'compound quality' advocate who told me that I should be taking a more drug-like malaria medication would be a two-fingered gesture that has occasionally been attributed to the English longbowmen at Agincourt. Some of the objection to quinones appears to be due to their ability to generate hydrogen peroxide via redox cycling and, when one makes this criticism of compounds, it is a good idea to at demonstrate that one is at least aware that hydrogen peroxide is an integral component of the regulatory mechanism of PTP1B.
This a good point to wrap things up. I just want to reiterate the importance of making a clear distinction in science between what you know and what you believe. This echoes Feynman who is reported to have said that, "The first principle is that you must not fool yourself and you are the easiest person to fool". Drug discovery is really difficult and I'm well aware that we often have to make decisions with incomplete information. When basing decisions on conclusions from data analysis, it is important to be fully aware of any assumptions that have been made and of limitations in what I'll call the 'scope' of the data. The output of forty high throughput screens that use different detection technologies is more likely to reveal genuinely pathological behavior than the output of six high throughput screens that all use the same detection technology. One needs to be very careful when extrapolating frequent hitter behavior to thiol reactivity and especially so when using results from a small number of assays that use a single detection technology. This article on antifungal PAINS is heavy on speculation (I counted 21 instances of 'could', 10 instances of 'might' and 5 instances of 'potentially') and light on evidence. I'm not denying that some (much?) of the output from high throughput screens is of minimal value and one key challenge is how to detect anc characterize bad chemical behavior in an objective manner. We need to think very carefully about how the 'PAINS' term should be used and the criteria by which compounds are classified as PAINS. Do we actually need to observe pan-assay interference in order to apply the term 'PAINS' to a compound or is it simply necessary for the compound to share a substructural feature with a minimum number of compounds for which pan-assay interference has been observed? How numerous and diverse must the assays be? The term 'PAINS' seems to get used more and more to describe any type of bad behavior (real, suspected or imagined) by compounds in assays and a case could be made for going back to talking about 'false positives' when referring to generic bad behavior in screens.
And I think that I'll leave it there. Hopefully provided some food for thought.
The authors present chemical structures of thirteen other compounds that they find unwholesome and I certainly wouldn't be be volunteering to optimize any of the compounds presented in this article. As an aside, when projects are handed over at transition time, it is instructive to observe how perceptions of compound quality differ between those trying to deliver a project and those charged with accepting it. My main criticism of the article is that very little evidence that bad things are actually happening is presented. The authors seem to be of the view that reactivity towards thiols is necessarily a Bad Thing. While formation of covalent bonds between ligands and proteins may be frowned upon in Switzerland (at least on Sundays), it remains a perfectly acceptable way to tame errant targets. A couple of quinones also appear in the rogues gallery and it needs to be pointed out that the naphthaquinone atovaquone is one of the two components (the other is proguanil which would also cause many compound quality 'experts' to spit feathers if they had failed to recognize it as an approved drug) of the antimalarial drug Malarone. I have actually taken Malarone on several occasions and the 'quinone-ness' of atovaquone worries me a great deal less than the potential neuropsychiatric effects of the (arguably) more drug-like mefloquine that is a potential alternative. My reaction to a 'compound quality' advocate who told me that I should be taking a more drug-like malaria medication would be a two-fingered gesture that has occasionally been attributed to the English longbowmen at Agincourt. Some of the objection to quinones appears to be due to their ability to generate hydrogen peroxide via redox cycling and, when one makes this criticism of compounds, it is a good idea to at demonstrate that one is at least aware that hydrogen peroxide is an integral component of the regulatory mechanism of PTP1B.
This a good point to wrap things up. I just want to reiterate the importance of making a clear distinction in science between what you know and what you believe. This echoes Feynman who is reported to have said that, "The first principle is that you must not fool yourself and you are the easiest person to fool". Drug discovery is really difficult and I'm well aware that we often have to make decisions with incomplete information. When basing decisions on conclusions from data analysis, it is important to be fully aware of any assumptions that have been made and of limitations in what I'll call the 'scope' of the data. The output of forty high throughput screens that use different detection technologies is more likely to reveal genuinely pathological behavior than the output of six high throughput screens that all use the same detection technology. One needs to be very careful when extrapolating frequent hitter behavior to thiol reactivity and especially so when using results from a small number of assays that use a single detection technology. This article on antifungal PAINS is heavy on speculation (I counted 21 instances of 'could', 10 instances of 'might' and 5 instances of 'potentially') and light on evidence. I'm not denying that some (much?) of the output from high throughput screens is of minimal value and one key challenge is how to detect anc characterize bad chemical behavior in an objective manner. We need to think very carefully about how the 'PAINS' term should be used and the criteria by which compounds are classified as PAINS. Do we actually need to observe pan-assay interference in order to apply the term 'PAINS' to a compound or is it simply necessary for the compound to share a substructural feature with a minimum number of compounds for which pan-assay interference has been observed? How numerous and diverse must the assays be? The term 'PAINS' seems to get used more and more to describe any type of bad behavior (real, suspected or imagined) by compounds in assays and a case could be made for going back to talking about 'false positives' when referring to generic bad behavior in screens.
And I think that I'll leave it there. Hopefully provided some food for thought.
No comments:
Post a Comment